Circulation Research 2021:
Clonal Hematopoiesis-Driver DNMT3A Mutations Alter Immune Cells in Heart Failure
Clonal hematopoiesis driven by mutations of DNMT3A (DNA methyltransferase 3a) is associated with increased incidence of cardiovascular disease and poor prognosis of patients with chronic heart failure (HF) and aortic stenosis. Although experimental studies suggest that DNMT3A clonal hematopoiesis-driver mutations may enhance inflammation, specific signatures of inflammatory cells in humans were missing. Wesley Abplanalp and his team at the Institute of Cardiovascular Regeneration used single cell sequencing to show, that heart failure patients with DNMT3A clonal hematopoiesis driver mutations show elevated inflammatory signatures in monocytes and T-cells at single cell resolution which may contribute to aggravation of heart failure.
Clonal hematopoiesis of indeterminate potential (CHIP) is an important CV risk factor
Clonal hematopoiesis is driven by acquired somatic mutations of regulatory genes in hematopoietic stem cells, leading to the clonal expansion of mutated blood cells. Clonal hematopoiesis is typically caused by myeloid leukemia-driver mutations, but only very few individuals carrying mutations associated with clonal hematopoiesis develop hematologic abnormalities. Therefore, the term “clonal hematopoiesis of indeterminate potential” (CHIP) was introduced. CHIP was shown to be associated with increased mortality, and the higher mortality of patients carrying CHIP-driver mutations was related to increased cardiovascular disease. More recently, CHIP emerged as an important cardiovascular risk factor, which contributes to the development of atherosclerosis and is associated with a poor prognosis of patients with heart failure and aortic stenosis. The most commonly mutated CHIP-driver genes include the DNA methyltransferase DNMT3A. Interestingly, in patients with aortic stenosis, Th17 to regulatory T-cell ratios were augmented in patients with DNMT3A mutations. Yet little information is available regarding potential pathways, which are activated or inactivated in monocyte and T-cell subpopulations in CHIP mutant carriers in humans.
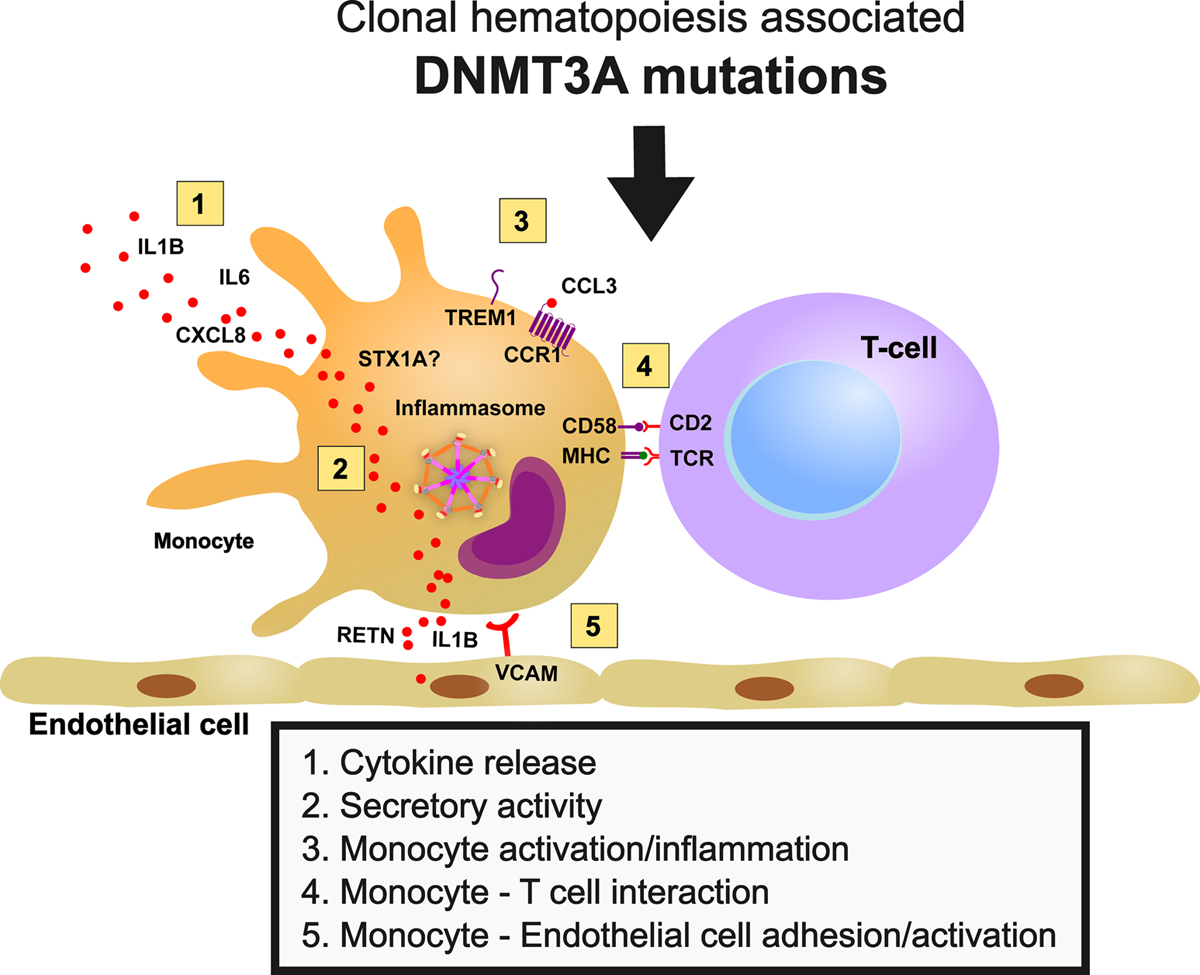
Figure: Schematics outlining influence of monocyte DNMT3A loss of function on the vascular milieu.
Single-cell RNA sequencing reveals inflamed transcriptome
In this study, Abplanalp et al. used single cell RNA sequencing to gain deeper insights into gene expression patterns, which might be associated with functional changes of circulating cells in patients with chronic heart failure (HF) harboring DNMT3A CHIP-driver mutations. Single-cell RNA sequencing (scRNA-seq) provides a unique and novel opportunity to define the transcriptome of individual cells, thereby disclosing transcriptional signatures in populations of cells. This study demonstrates that circulating monocytes and T cells of patients with HF harboring clonal hematopoiesis-driver mutations in DNMT3A exhibit a highly inflamed transcriptome, which may contribute to the aggravation of chronic HF.
Systemic anti-inflammatory strategy as therapeutic option
Knowing this critical information could enable insights as to how to specifically block pathological inflammation in a tailored manner or by a systemic anti-inflammatory strategy, such as anti-IL-1B therapy as has been used in the CANTOS trial.
Find the original publication here...
There is also commentary on the article through the links below:
https://www.ahajournals.org/doi/10.1161/CIRCRESAHA.120.318575
https://www.nature.com/articles/s41569-020-00481-5
An interview with the authors can be found here:
https://circresearch.libsyn.com/january-2021-discover-circres
